nLab Introduction to Spectral Sequences
This page is an introduction to spectral sequences. We motivate spectral sequences of filtered complexes from the computation of cellular cohomology via stratum-wise relative cohomology. In the end we generalize to spectral sequences of filtered spectra.
For background on homological algebra see at Introduction to Homological algebra.
For background on stable homotopy theory see at Introduction to Stable homotopy theory.
For application to complex oriented cohomology see at Introduction to Cobordism and Complex Oriented Cohomology.
For application to the Adams spectral sequence see Introduction to Adams spectral sequences.
Contents
In Introduction to Stable homotopy theory we have set up the concept of spectra and their stable homotopy groups (def.). More generally for and two spectra then there is the graded stable homotopy group of homotopy classes of maps bewteen them (def.). These may be thought of as generalized cohomology groups (exmpl.). Moreover, in part 1.2 we discussed the symmetric monoidal smash product of spectra . The stable homotopy groups of such a smash product spectrum may be thought of as generalized homology groups (rmk.).
These stable homotopy and generalized (co-)homology groups are the fundamental invariants in algebraic topology. In general they are as rich and interesting as they are hard to compute, as famously witnessed by the stable homotopy groups of spheres, some of which we compute in part 2.
In general the only practicable way to carry out such computations is by doing them along a decomposition of the given spectrum into a “sequence of stages” of sorts. The concept of spectral sequence is what formalizes this idea.
(Here the re-occurence of the root “spectr-” it is a historical coincidence, but a lucky one.)
Here we give an expository introduction to the concept of spectral sequences, building up in detail to the spectral sequence of a filtered complex.
We put these spectral sequences to use in
For filtered complexes
We begin with recalling basics of ordinary relative homology and then seamlessly derive the notion of spectral sequences from that as the natural way of computing the ordinary cohomology of a CW-complex stagewise from the relative cohomology of its skeleta. This is meant as motivation and warmup. What we are mostly going to use further below are spectral sequences induced by filtered spectra, this we turn to next.
Ordinary homology
Let be a topological space and a topological subspace. Write for the chain complex of singular homology on and for the chain map induced by the subspace inclusion.
Definition
The (degreewise) cokernel of this inclusion, hence the quotient of by the image of under the inclusion, is the chain complex of -relative singular chains.
-
A boundary in this quotient is called an -relative singular boundary,
-
a cycle is called an -relative singular cycle.
-
The chain homology of the quotient is the -relative singular homology of
Remark
This means that a singular -chain is an -relative cycle precisely if its boundary is, while not necessarily 0, contained in the -chains of : . So the boundary vanishes possibly only “up to contributions coming from ”.
We record two evident but important classes of long exact sequences that relative homology groups sit in:
Proposition
Let be a topological subspace inclusion. The corresponding relative singular homology, def. , sits in a long exact sequence of the form
The connecting homomorphism sends an element represented by an -relative cycle , to the class represented by the boundary .
Proof
This is the homology long exact sequence, induced by the defining short exact sequence of chain complexes.
Proposition
Let be a sequence of two topological subspace inclusions. Then there is a long exact sequence of relative singular homology groups of the form
Proof
Observe that we have a short exact sequence of chain complexes, def.
The corresponding homology long exact sequence, prop. , is the long exact sequence in question.
We look at some concrete fundamental examples in a moment. But first it is useful to make explicit the following general sub-notion of relative homology.
Let still be a given topological space.
Definition
The augmentation map for the singular homology of is the homomorphism of abelian groups
which adds up all the coefficients of all 0-chains:
Since the boundary of a 1-chain is in the kernel of this map, by example , it constitutes a chain map
where now is regarded as a chain complex concentrated in degree 0.
Definition
The reduced singular chain complex of is the kernel of the augmentation map, the chain complex sitting in the short exact sequence
The reduced singular homology of is the chain homology of the reduced singular chain complex
Equivalently:
Definition
The reduced singular homology of , denoted , is the chain homology of the augmented chain complex
Let be a topological space, its singular homology and its reduced singular homology, def. .
Proposition
For there is an isomorphism
Proof
The homology long exact sequence, prop. , of the defining short exact sequence is, since here is concentrated in degree 0, of the form
Here exactness says that all the morphisms for positive are isomorphisms. Moreover, since is a free abelian group, hence a projective object, the remaining short exact sequence
is split, by prop. , and hence .
Proposition
For the point, the morphism
is an isomorphism. Accordingly the reduced homology of the point vanishes in every degree:
Now we can discuss the relation between reduced homology and relative homology.
Proposition
For an inhabited topological space, its reduced singular homology, def. , coincides with its relative singular homology relative to any base point :
Proof
Consider the sequence of topological subspace inclusions
By prop. this induces a long exact sequence of the form
Here in positive degrees we have and therefore exactness gives isomorphisms
and hence with prop. isomorphisms
It remains to deal with the case in degree 0. To that end, observe that is a monomorphism: for this notice that we have a commuting diagram
where is the terminal map. That the outer square commutes means that and hence the composite on the left is an isomorphism. This implies that is an injection.
Therefore we have a short exact sequence as shown in the top of this diagram
Using this we finally compute
With this understanding of homology relative to a point in hand, we can now characterize relative homology more generally. From its definition in def. , it is plausible that the relative homology group provides information about the quotient topological space . This is indeed true under mild conditions:
Definition
A topological subspace inclusion is called a good pair if
-
is closed inside ;
-
has an neighbourhood such that has a deformation retract.
Proposition
If is a topological subspace inclusion which is good in the sense of def. , then the -relative singular homology of coincides with the reduced singular homology, def. , of the quotient space :
The proof of this is spelled out at Relative homology – relation to quotient topological spaces. It needs the proof of the Excision property of relative homology. While important, here we will not further dwell on this. The interested reader can find more information behind the above links.
Cellular homology
With the general definition of relative homology in hand, we now consider the basic cells such that cell complexes built from such cells have tractable relative homology groups. Actually, up to weak homotopy equivalence, every Hausdorff topological space is given by such a cell complex and hence its relative homology, then called cellular homology, is a good tool for computing singular homology rather generally.
Definition
For write
Example
The reduced singular homology of the -sphere equals the -relative homology of the -disk with respect to the canonical boundary inclusion : for all
Proof
The -sphere is homeomorphic to the -disk with its entire boundary identified with a point:
Moreover the boundary inclusion is a good pair in the sense of def. . Therefore the example follows with prop. .
When forming cell complexes from disks, then each relative dimension will be a wedge sum of disks:
Definition
For a set of pointed topological spaces, their wedge sum is the result of identifying all base points in their disjoint union, hence the quotient
Example
The wedge sum of two pointed circles is the “figure 8”-topological space.
Proposition
Let be a set of pointed topological spaces. Write for their wedge sum and write for the canonical inclusion functions.
Then for each the homomorphism
is an isomorphism.
Proof
By prop. the reduced homology of the wedge sum is equivalently the relative homology of the disjoint union of spaces relative to their disjoint union of basepoints
The relative homology preserves these coproducts (sends them to direct sums) and so
The following defines topological spaces which are inductively built by gluing disks to each other.
Definition
A CW complex of dimension is the empty topological space.
By induction, for a CW complex of dimension is a topological space obtained from
-
a -complex of dimension ;
-
an index set ;
-
a set of continuous maps (the attaching maps)
as the pushout
in
hence as the topological space obtained from by gluing in -disks for each along the given boundary inclusion .
By this construction, an -dimensional CW-complex is canonically a filtered topological space, hence a sequence of topological subspace inclusions of the form
which are the right vertical morphisms in the above pushout diagrams.
A general CW complex then is a topological space which is the limiting space of a possibly infinite such sequence, hence a topological space given as the sequential colimit over a tower diagram each of whose morphisms is such a filter inclusion
The following basic facts about the singular homology of CW complexes are important.
Now we can state a variant of singular homology adapted to CW complexes which admits a more systematic way of computing its homology groups. First we observe the following.
Proposition
The relative singular homology, def. , of the filtering degrees of a CW complex , def. , is
where denotes the free abelian group on the set of -cells.
Proof
The inclusion is a good pair in the sense of def. . The quotient is by definition of CW-complexes a wedge sum, def. , of -spheres, one for each element in . Therefore by prop. we have an isomorphism with the reduced homology of this wedge sum. The statement then follows by the respect of reduced homology for wedge sums, prop. .
Proposition
For a CW complex with skeletal filtration as above, and with we have for the singular homology of that
In particular if is a CW-complex of finite dimension (the maximum degree of cells), then
Moreover, for the inclusion
is an isomorphism and for we have an isomorphism
Proof
By the long exact sequence in relative homology, prop. we have an exact sequence of the form
Now by prop. the leftmost and rightmost homology groups here vanish when and and hence exactness implies that
is an isomorphism for . This implies the first claims by induction on .
Finally for the last claim use that the above exact sequence gives
and hence that with the above the map is surjective.
We may now discuss the cellular homology of a CW complex.
Definition
For a CW-complex, def. , its cellular chain complex is the chain complex such that for
-
the abelian group of chains is the relative singular homology group, def. , of relative to :
-
the differential is the composition
where is the boundary map of the singular chain complex and where is the morphism on relative homology induced from the canonical inclusion of pairs .
Proposition
The composition of two differentials in def. is indeed zero, hence is indeed a chain complex.
Proof
On representative singular chains the morphism acts as the identity and hence acts as the double singular boundary, .
Remark
This means that
-
a cellular -chain is a singular -chain required to sit in filtering degree , hence in ;
-
a cellular -cycle is a singular -chain whose singular boundary is not necessarily 0, but is contained in filtering degree , hence in .
-
a cellular -boundary is a singular -chain which is the boundary of a singular -chain coming from filtering degree .
This kind of situation – chains that are cycles only up to lower filtering degree and boundaries that come from specified higher filtering degree – has an evident generalization to higher relative filtering degrees. And in this greater generality the concept is of great practical relevance. Therefore before discussing cellular homology further now, we consider this more general “higher-order relative homology” that it suggests (namely the formalism of spectral sequences). After establishing a few fundamental facts about that we will come back in prop. below to analyse the above cellular situation using this conceptual tool.
In theorem we conclude that cellular homology and singular homology agree of CW-complexes agres.
First we abstract the structure on chain complexes that in the above example was induced by the CW-complex structure on the singular chain complex.
Filtered chain complexes
Definition
The structure of a filtered chain complex in a chain complex is a sequence of chain map inclusions
The associated graded complex of a filtered chain complex, denoted , is the collection of quotient chain complexes
We say that element of are in filtering degree .
Remark
In more detail this means that
-
is a chain complex, hence are objects in (-modules) and are morphisms (module homomorphisms) with ;
-
For each there is a filtering on and all these filterings are compatible with the differentials in that
-
The grading associated to the filtering is such that the -graded elements are those in the quotient
Since the differentials respect the grading we have chain complexes in each filtering degree .
Hence elements in a filtered chain complex are bi-graded: they carry a degree as elements of as usual, but now they also carry a filtering degree: for we therefore also write
and call this the collection of -chains in the filtered chain complex.
Accordingly we have -cycles and -boundaries. But for these we may furthermore refine to a notion where also the filtering degree of the boundaries is constrained:
Definition
Let be a filtered chain complex. Its associated graded chain complex is the set of chain complexes
for all .
Then for we say that
-
is the module of -chains or of -chains in filtering degree ;
-
the module
is the module of -almost -cycles (the -chains whose differential vanishes modulo terms of filtering degree );
-
is the module of -almost -boundaries.
Similarly we set
From this definition we immediately have that the differentials restrict to the -almost cycles as follows:
Proposition
The differentials of restrict on -almost cycles to homomorphisms of the form
These are still differentials: .
Proof
By the very definition of it consists of elements in filtering degree on which decreases the filtering degree to . Also by definition of differential on a chain complex, decreases the actual degree by one. This explains that restricted to lands in . Now the image constists indeed of actual boundaries, not just -almost boundaries. But since actual boundaries are in particular -almost boundaries, we may take the codomain to be .
As before, we will in general index these differentials by their codomain and hence write in more detail
Proposition
We have a sequence of canonical inclusions
The following observation is elementary, and yet this is what drives the theory of spectral sequences, as it shows that almost cycles may be computed iteratively by homological means themselves.
Proposition
The -almost cycles are the -kernel inside the -almost cycles:
Proof
An element represents
-
an element in if
-
an element in if even .
The second condition is equivalent to representing the 0-element in the quotient . But this is in turn equivalent to being 0 in .
With a definition of almost-cycles and almost-boundaries, of course we are now interested in the corresponding homology groups:
Definition
For define the -almost -chain homology of the filtered complex to be the quotient of the -almost -cycles by the -almost -boundaries, def. :
By prop. the differentials of restrict on the -almost homology groups to maps
The central property of these -almost homology groups now is their following iterative homological characterization.
Proposition
With definition we have that is the -chain homology of :
This structure on the collection of -almost cycles of a filtered chain complex thus obtained is called a spectral sequence:
Definition
A homology spectral sequence of -modules is
-
a set of -modules;
-
a set of homomorphisms
such that
-
the s are differentials: ;
-
the modules are the -homology of the modules in relative degree :
One says that is the -page of the spectral sequence.
Since this turns out to be a useful structure to make explicit, as the above motivation should already indicate, one introduces the following terminology and basic facts to talk about spectral sequences.
Definition
Let be a spectral sequence, def. , such that for each there is such that for all we have
Then one says that
-
the bigraded object
is the limit term of the spectral sequence;
- the spectral sequence abuts to .
Example
If for a spectral sequence there is such that all differentials on pages after vanish, , then is a limit term for the spectral sequence. One says in this cases that the spectral sequence degenerates at .
Proof
By the defining relation
the spectral sequence becomes constant in from on if all the differentials vanish, so that for all .
Example
If for a spectral sequence there is such that the th page is concentrated in a single row or a single column, then the spectral sequence degenerates on this pages, example , hence this page is a limit term, def. . One says in this case that the spectral sequence collapses on this page.
Proof
For the differentials of the spectral sequence
have domain and codomain necessarily in different rows an columns (while for both are in the same row and for both coincide). Therefore if all but one row or column vanish, then all these differentials vanish.
Definition
A spectral sequence is said to converge to a graded object with filtering , traditionally denoted
if the associated graded complex of is the limit term of , def. :
Remark
In practice spectral sequences are often referred to via their first non-trivial page, often also the page at which it collapses, def. , often already the second page. Then one tends to use notation such as
to be read as “There is a spectral sequence whose second page is as shown on the left and which converges to a filtered object as shown on the right.”
Definition
A spectral sequence is called a bounded spectral sequence if for all the number of non-vanishing terms of total degree , hence of the form , is finite.
Definition
A spectral sequence is called
-
a first quadrant spectral sequence if all terms except possibly for vanish;
-
a third quadrant spectral sequence if all terms except possibly for vanish.
Proof
First notice that if a spectral sequence has at most non-vanishing terms of total degree on page , then all the following pages have at most at these positions non-vanishing terms, too, since these are the homologies of the previous terms.
Therefore for a bounded spectral sequence for each there is such that for all and all . Similarly there is such for all and all .
We claim then that the limit term of the bounded spectral sequence is in position given by the value for
This is because for such we have
Therefore
The central statement about the notion of the spectral sequence of a filtered chain complex then is the following proposition. It says that the iterative computation of higher order relative homology indeed in the limit computes the genuine homology.
Definition
For a filtered complex, write for
This defines a filtering of the homology, regarded as a graded object.
Proposition
If the spectral sequence of a filtered complex of prop. has a limit term, def. then it converges, def. , to the chain homology of
i.e. for sufficiently large we have
where on the right we have the associated graded object of the filtering of def. .
Proof
By assumption, there is for each an such that for all the -almost cycles and -almost boundaries, def. , in are the ordinary cycles and boundaries. Therefore for def. gives .
This says what these spectral sequences are converging to. For computations it is also important to know how they start out for low . We can generally characterize for very low values of simply as follows:
Proof
because for we automatically also have since the differential respects the filtering degree by assumption.
Remark
There is, in general, a decisive difference between the homology of the associated graded complex and the associated graded piece of the genuine homology : in the former the differentials of cycles are required to vanish only up to terms in lower degree, but in the latter they are required to vanish genuinely. The latter expression is instead the value of the spectral sequence for , see prop. below.
Comparing cellular and singular homology
These general facts now allow us, as a first simple example for the application of spectral sequences to see transparently that the cellular homology of a CW complex, def. , coincides with its genuine singular homology.
First notice that of course the structure of a CW-complex on a topological space , def. naturally induces on its singular simplicial complex the structure of a filtered chain complex, def. :
Definition
For a CW complex, and , write
for the singular chain complex of . The given topological subspace inclusions induce chain map inclusions and these equip the singular chain complex of with the structure of a bounded filtered chain complex
(If is of finite dimension then this is a bounded filtration.)
Write for the spectral sequence of a filtered complex corresponding to this filtering.
Proposition
The spectral sequence of singular chains in a CW complex , def. converges, def. , to the singular homology of :
Proof
The spectral sequence is clearly a first-quadrant spectral sequence, def. . Therefore it is a bounded spectral sequence, def. and hence has a limit term, def. . So the statement follows with prop. .
We now identify the low-degree pages of with structures in singular homology theory.
Proposition
-
– is the group of -relative (p+q)-chains, def. , in ;
-
– is the -relative singular homology, def. , of ;
-
–
-
– .
Proof
By straightforward and immediate analysis of the definitions.
As a result of these general considerations we now obtain the promised isomorphism between the cellular homology and the singular homology of a CW-complex :
Theorem
For Top a CW complex, def. , its cellular homology, def. coincides with its singular homology :
Proof
By the third item of prop. the -page of the spectral sequence is concentrated in the -row and hence it collapses there, def. . Accordingly we have
for all . By the third and fourth item of prop. this non-trivial only for and there it is equivalently
Finally observe that by the definition of the filtering on the homology, def. , and using prop. .
For filtered spectra
Definition
A filtered spectrum is a spectrum equipped with a sequence of spectra of the form
Remark
More generally a filtering on an object in (stable or not) homotopy theory is a -graded sequence such that is the homotopy colimit . But for the present purpose we stick with the simpler special case of def. .
Remark
There is no condition on the morphisms in def. . In particular, they are not required to be n-monomorphisms or n-epimorphisms for any .
On the other hand, while they are also not explicitly required to have a presentation by cofibrations or fibrations, this follows automatically: by the existence of model structures for spectra, every filtering on a spectrum is equivalent to one in which all morphisms are represented by cofibrations or by fibrations.
This means that we may think of a filtration on a spectrum in the sense of def. as equivalently being a tower of fibrations over .
The following remark unravels the structure encoded in a filtration on a spectrum, and motivates the concepts of exact couples and their spectral sequences from these.
Remark
Given a filtered spectrum as in def. , write for the homotopy cofiber of its th stage, such as to obtain the diagram
where each stage
is a homotopy fiber sequence.
To break this down into invariants, apply the stable homotopy groups-functor (def.). This yields a diagram of -graded abelian groups of the form
Each hook at stage extends to a long exact sequence of homotopy groups (prop.) via connecting homomorphisms
If we understand the connecting homomorphism
as a morphism of degree -1, then all this information fits into one diagram of the form
where each triangle is a rolled-up incarnation of a long exact sequence of homotopy groups (and in particular is not a commuting diagram!).
If we furthermore consider the bigraded abelian groups and , then this information may further be rolled-up to a single diagram of the form
where the morphisms , and have bi-degree , and , respectively.
Here it is convenient to shift the bigrading, equivalently, by setting
because then counts the cycles of going around the triangles:
Data of this form is called an exact couple, def. below.
Definition
An unrolled exact couple (of Adams-type) is a diagram of abelian groups of the form
such that each triangle is a rolled-up long exact sequence of abelian groups of the form
The collection of this “un-rolled” data into a single diagram of abelian groups is called the corresponding exact couple.
Definition
An exact couple is a diagram (non-commuting) of abelian groups of the form
such that this is exact sequence exact in each position, hence such that the kernel of every morphism is the image of the preceding one.
The concept of exact couple so far just collects the sequences of long exact sequences given by a filtration. Next we turn to extracting information from this sequence of sequences.
Remark
The sequence of long exact sequences in remark is inter-locking, in that every appears in two of them, and thus we can string them all together:
This gives rise to the horizontal composites , as show above, and by the fact that the diagonal sequences are long exact, these are differentials: , hence give a chain complex:
We read off from the interlocking long exact sequences what these differentials mean: an element lifts to an element precisely if :
This means that the cochain cohomology of the complex produces elements of and hence of .
In order to organize this observation, notice that in terms of the exact couple of remark , the differential
is a component of the composite
Some terminology:
Definition
Given an exact couple, def. , then the induced derived exact couple is the diagram
with
-
;
-
;
-
;
-
;
-
.
Definition
Given an exact couple, def. , then the induced spectral sequence, def. , is the sequence of pages, def. , of the induced sequence of derived exact couples, def. , prop. .
Example
Consider a filtered spectrum, def. ,
and its induced exact couple of stable homotopy groups, from remark
with bigrading as shown on the right.
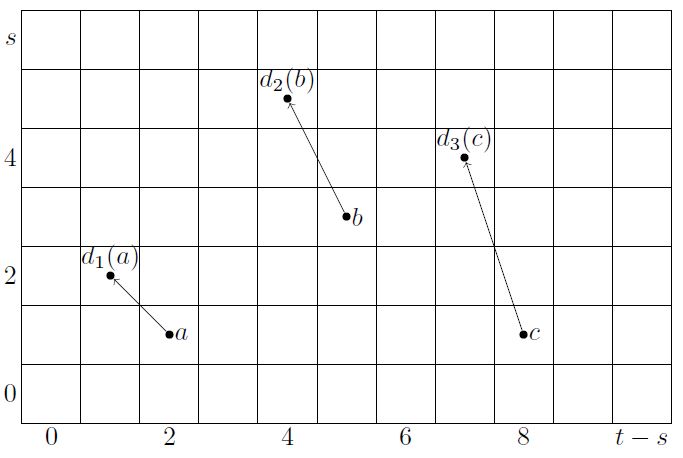
As we pass to derived exact couples, by def. , the bidegree of and is preserved, but that of increases by in each step, since
Therefore the induced spectral sequence has differentials of the form
This is also called the Adams-type spectral sequence of the tower of fibrations .
This we discuss in detail in part 2 – Adams spectral sequences.
References
A gentle exposition of the general idea of spectral sequences is in
- John McCleary, A User’s Guide to Spectral Sequences, Cambridge University Press (1985, 2001)
A concise account streamlined for our purposes is in section 2 of
- John Rognes, The Adams spectral sequence (following Bruner), 2012 (pdf)
Last revised on December 5, 2021 at 06:28:36. See the history of this page for a list of all contributions to it.